Understanding the alphabet soup of Coronavirus SARS-CoV-2 genome. B.1.1.7, Delta, E484K… What do the numbers (and letters) mean?
News reports on Covid and the virus responsible for the infection, SARS-CoV-2, is on the news with plenty of science-speak numbers and acronyms. But because these acronyms are usually not part of the ‘news’ being covered in the story, exactly what those codes mean is rarely explained. For example, former FDA Commissioner Scott Gottlieb recently tweeted:
“UK reported its biggest one-day Covid case increase in 3 months just as the new delta variant AY.4 with the S:Y145H mutation in the spike reaches 8% of UK sequenced cases. We need urgent research to figure out if this delta plus is more transmissible, has partial immune evasion?
Delta, AY.4, S:Y145H from Gottlieb’s tweet are all opaque to most readers, (we presume they know UK refers to United Kingdom…), and clearly, the urgency of this tweet demanded attention be focused on the spread and virulence of the then-new worrisome strain (“Delta Plus”) rather than a mundane explanation of the acronyms. The delta plus strain includes the K417N mutation, which may increase the virus strain’s ability to reinfect survivors of earlier Covid strain infections.
and
“…a delta sublineage newly designated as AY.4.2 is noted to be expanding in England… This sublineage is currently increasing in frequency. It includes spike mutations A222V and Y145H." (https://www.cnbc.com/2021/10/20/uk-doctors-call-for-return-of-covid-restrictions-new-mutation-watched.html)
SARS-CoV-2 genome
The genome of the coronavirus responsible for Covid, technically called SAS-CoV-2, is a ring of RNA almost 30,000 bases long. The bases are like a long string of letters spelling words, but in this case, RNA uses only 4 letters (A,U,C, and G) and the words spelled out by the exact sequence of letters in structural genes are instructions for the infected human cell to make specific kinds of proteins. In the structural gene "coding" portion of the genome, each three RNA bases, called a triplet, is like a word calling for a specific amino acid. For example, if the RNA triplet base sequence is AUG, that calls for the amino acid methionine, which is added to the growing chain of amino acids that eventually becomes a functional protein.
The SARS-CoV-2 genome spells out about two dozen different proteins, many of which are important in the infection process or the severity of the resulting illness.
One example is the “spike” protein, which sticks out of the virus particle like points on a crown (and hence the name ‘coronavirus’). The spike protein is responsible for the virus docking with their target human cells, especially the ACE-2 cells) to start the infection process. The Spike protein is also the target for current mRNA based vaccines.
Mutations are permanent changes to the base sequence of letters in the genome, whether the genome is RNA (like coronavirus and many other viruses) or DNA (like other viruses, and all other living things, including humans). Mutations happen in all living things and can be induced by certain chemicals, environmental conditions, or just spontaneously. Mutations can affect a single base, or several bases, or even very long stretches of bases, and can involve deletion of the base(s), or changing a base from one letter to another, or duplication of the base(s), or inversion, insertion of additional bases, or virtually anything that can go wrong with the genome.
Many known mutations in SARS-CoV-2 are single base changes, called point mutations. Because the viral genome is short compared to other living things, the mutation of a large number of bases will likely render the virus weak or incompetent, effectively killing it off as a pathogen. Any quickly growing population (not just viruses) will tend to generate more mutations, and the bigger the population, the more mutations will accumulate. At least some of these mutations will likely make the pathogen even more virulent.
While this mutation stuff is fascinating for molecular geneticists like me, most people just want to know how they relate to Covid. Well here it is— each mutation has the potential to increase infectivity, or severity of illness, or evade antibodies, so scientific management of Covid requires knowing each one precisely, which requires providing names. Each new variant or strain carries one or more mutations in the genome. The naming convention started haphazardly, based on the lab which initially identified and characterized each mutation, or the geographic source where the strain was first identified. As identification and documentation of variants increased this practice became unwieldy and prone to ambiguity and misinterpretation, so WHO (World Health Organization) took over to standardize the naming procedure, using Greek letters.
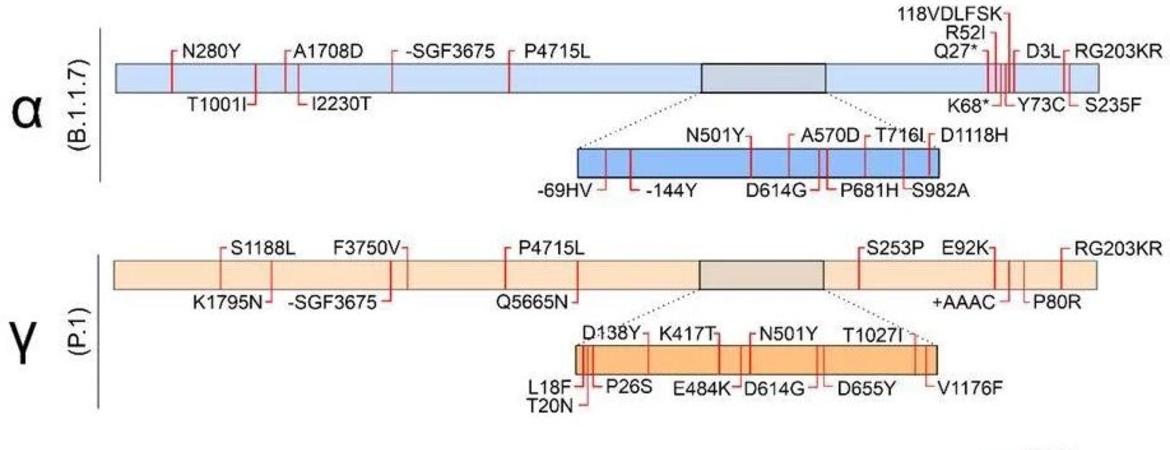
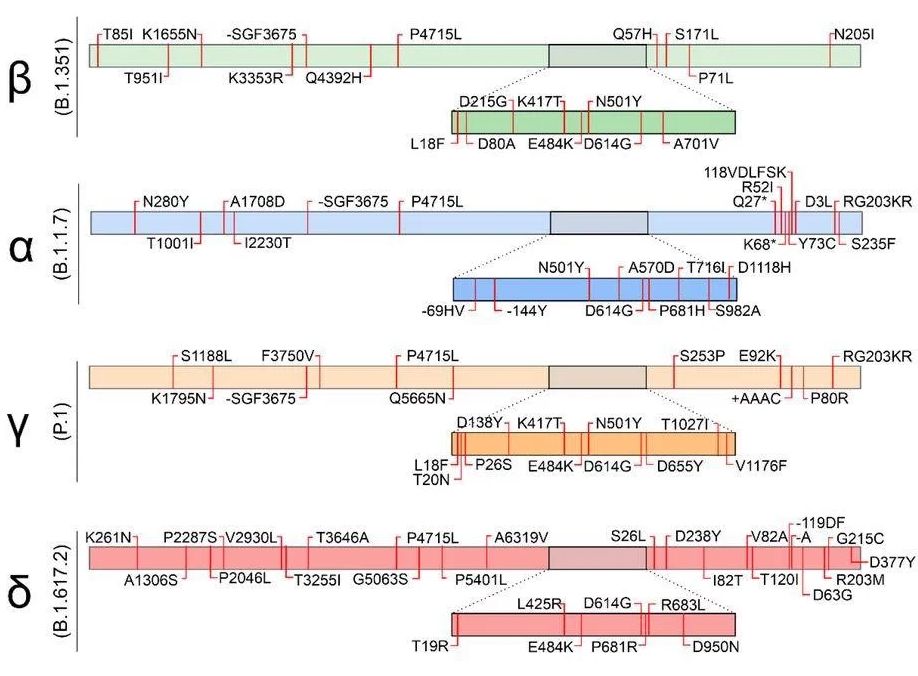
Under WHO, the identity of the first group of “variants of concern” (i.e., those strains with potentially increased pathogenicity, either in infectivity or increased severity of illness potential) went from B.1.1.7 to Alpha, B1.351 to Beta, P.1 to Gamma and B.1.617.2 to Delta, the first four letters of the Greek alphabet. Omicron (also known as B.1.1.529 under the old nomenclature system) is the 5th “Variant of Concern” and undoubtedly will go well beyond.
Each Greek letter refers to a known variant strain with identified mutations in the genome. Each variant of concern has multiple mutations in their RNA sequence, and specifically in sequences providing the recipes for Spike and other structural proteins involved in pathogenicity. That is, mutations in the gene for the spike protein can lead to a strain with spike proteins better able to dock with human target cells (and thus increase infectivity) or spike proteins less susceptible to mRNA vaccine antibodies that recognize specific parts of the spike protein. Omicron, for example, has over 30 mutations in the spike protein gene alone. Note, however, that the absolute number of mutations does not correlate with altered pathogenicity. A single mutation at a particular location on the gene converting one specific amino acid might have more impact than 50 mutations scattered elsewhere. Omicron carries several mutations, including base substitutions resulting in amino acid substitutions, base deletions, base insertions, and more. Some of these are especially worrisome because they are known to influence pathogenicity.
Scientists need a means to identify those specific mutations within the Greek letter strain designation. The exact amino acid sequence of a given protein is determined by the exact base sequence of the respective gene. If a mutation in the gene's RNA sequence changes, say, converting a G to an A, the resulting protein might have a crucial amino acid change. So, for example. E484K refers to the Spike protein’s 484th amino acid in the protein chain. The amino acid glutamine (abbreviated ‘E’) at that exact location in the ancestral strain is replaced by the amino acid Lysine (abbr. ‘K’). This apparently small difference of one amino acid in the spike protein can have grave consequences, as this mutation seems to help the virus evade immune responses, including the protective effect of vaccines. Importantly, this E484K mutation is found in Beta, Gamma and Delta. Each mutation causing a change in the amino acid sequence of proteins involved in pathogenicity must be carefully scrutinized in each new strain as they arise. The best defense against new, potentially pathogenic mutations is to limit the population of viral particles. That is best achieved by vaccinations, masking and social distancing.
More info:
https://www.newsweek.com/delta-ay-variant-ay4-2-urgent-research-scott-gottlieb-fda-which-us-states-uk-1639842
https://www.ncbi.nlm.nih.gov/gene/43740568
https://www.nytimes.com/interactive/2021/health/coronavirus-variant-tracker.html
https://www.newsweek.com/delta-ay-variant-ay4-2-urgent-research-scott-gottlieb-fda-which-us-states-uk-1639842
https://www.cdc.gov/coronavirus/2019-ncov/variants/delta-variant.html?s_cid=11512:covid%20delta:sem.ga:p:RG:GM:gen:PTN:FY21
https://www.sfchronicle.com/health/article/What-we-know-about-AY-4-2-a-delta-subvariant-16544024.php
https://pubmed.ncbi.nlm.nih.gov/33532796/ (E484K mutation)
https://en.wikipedia.org/wiki/Variants_of_SARS-CoV-2
https://www.bmj.com/content/372/bmj.n359 (E484K variant impact on immune response)
https://www.sciencealert.com/how-concerned-should-you-be-about-ay-4-2-delta-subvariant-an-expert-explains (AY-4-2 variant)
https://www.news-medical.net/news/20211122/Interferon-beta-versus-the-SARS-CoV-2-Delta-variant.aspx (Figure 2)
https://covariants.org/variants/21K.Omicron